Appendix B. List with the assessment of intrinsic identifiability of different temporary emigration models.
A pdf-version of Appendix B is also available for viewing.
Intrinsic identifiability
of the temporary emigration models, assessed with computer algebra (see Appendix
A for MAPLE code used for the assessment). Table entries
are the parameters that are separately identifiable, the number of estimated
parameters when k capture occasions are considered. Model notations follow
Lebreton et al. (1992). S: survival probability, ![]() :
transition probability between observable and unobservable state (emigration),
:
transition probability between observable and unobservable state (emigration),
![]() : transition
probability between unobservable and observable state (immigration), p:
on-site recapture probability. In the model notation, subscript t refers
to time, g to group, and . to constancy. In the parameter notation, superscript
g1 refers to group1, superscript g2 to group2, the subscript m
refers to the last capture occasion and subscript 1 to the first capture
occasion. Note:
: transition
probability between unobservable and observable state (immigration), p:
on-site recapture probability. In the model notation, subscript t refers
to time, g to group, and . to constancy. In the parameter notation, superscript
g1 refers to group1, superscript g2 to group2, the subscript m
refers to the last capture occasion and subscript 1 to the first capture
occasion. Note: ![]() is
never estimable, but this is not stated explicitly in this table.
is
never estimable, but this is not stated explicitly in this table.
Model
Identifiable parameters
Number of estimated quantities
1:
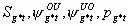
None
8k-20
2:
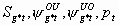
None
7k-18
3:
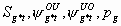
None, but pg1 and pg2
6k-10
4:

None, but p
6k-11
5:
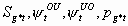
None, but pg12 and pg22
6k-13
6:
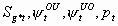
None, but p2
5k-11
7:
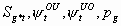
None, but pg1 and pg2
4k-5
8:
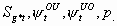
None, but p
4k-6
9:

All, but Sg1m, Sg2m, pg1m, pg2m
4k-2
10:
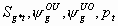
All, but Sg1m, Sg2m, pm
3k
11:
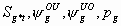
All
2k+4
12:

All
2k+3
13:
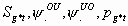
All, but Sg1m, Sg2m, pg1m, pg2m
4k-4
14:

All, but Sg1m, Sg2m, pm
3k-2
15:
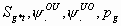
All
2k+2
16:
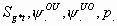
All
2k+1
17:
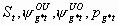
All, but Sm-1, Sm,
,
,
,
,
,
,
,
,
,
, p2, pm
7k-15
18:
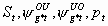
All, but Sm,
,
,
,
,
,
,
,
, p1, pm
6k-11
19:
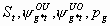
All, but Sm,
,
,
,
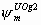
5k-6
20:
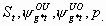
All, but Sm,
,
,
,
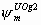
5k-7
21:
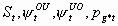
None, but pg12 and pg22
5k-11
22:
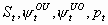
None
4k-10
23:
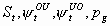
None, but pg1 and pg2
3k-4
24:
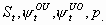
None, but p
3k-5
25:
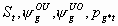
All, but Sm, pg1m, pg1m
3k
26:
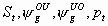
All, but Sm, pm
2k+1
27:
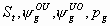
All
k+5
28:
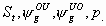
All
k+4
29:
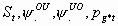
All, but Sm, pg1m, pg2m
3k-2
30:
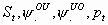
All, but Sm, pm
2k-1
31:
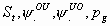
All
k+3
32:
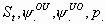
All
k+2
33:
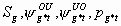
All, but
,
,
,
,
,
,
,
, pg11, pg21, pg1m, pg2m
6k-10
34:
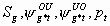
All, but
,
,
,
,
,
,
,
, p1, pm
5k-7
35:
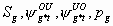
All
4k-2
36:
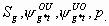
All
4k-3
37:
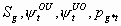
All, but
,
,
,
, pg11, pg1m, pg21, pg2m
4k-5
38:
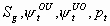
All, but
,
,
,
, p1, pm
3k-4
39:
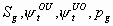
All
2k+1
40:
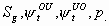
All
2k
41:
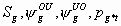
All
2k+4
42:
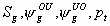
All
k+5
43:
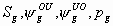
All
8
44:
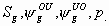
All
7
45:
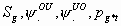
All
2k+2
46:
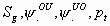
All
k+3
47:
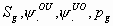
All
6
48:
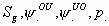
All
5
49:
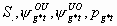
All, but
,
,
,
,
,
,
,
, pg11, pg21, pg1m, pg2m
6k-11
50:
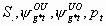
All but
,
,
,
,
,
,
,
, p1, pm
5k-8
51:
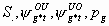
All
4k-3
52:
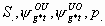
All
4k-4
53:
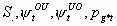
All, but
,
,
,
, pg11, pg1m, pg21, pg2m
4k-6
54:
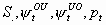
All, but
,
,
,
, p1, pm
3k-5
55:
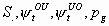
All
2k
56:
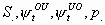
All
2k-1
57:
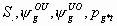
All
2k+3
58:
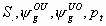
All
k+4
59:
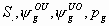
All
7
60:

All
6
61:
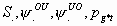
All
2k+1
62:
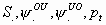
All
k+2
63:
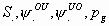
All
5
64:
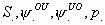
All
4
Literature cited
Lebreton, J.-D., K. P. Burnham, J. Clobert, and D. R. Anderson. 1992. Modelling survival and testing biological hypotheses using marked animals: a unified approach with case studies. Ecological Monographs 62:67–118.